BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:
URL: http://journal.zums.ac.ir/article-1-7238-en.html


 , Zeinab Dehghan2
, Zeinab Dehghan2 

 , Abbas Alibakhshi3
, Abbas Alibakhshi3 

 , Fatemeh Yarian4
, Fatemeh Yarian4 

 , Hakimeh Zali *5
, Hakimeh Zali *5 


2- Dept. of Comparative Biomedical Sciences, School of Advanced Medical Sciences and Technologies, Shiraz University of Medical Sciences, Shiraz, Iran
3- Molecular Medicine Research Center, Hamadan University of Medical Sciences, Hamadan, Iran
4- Dept. of Medical Biotechnology, School of Advanced Technologies in Medicine, Fasa University of Medical Sciences, Fasa, Iran
5- Dept. of Tissue Engineering and Applied Cell Sciences, School of Advanced Technologies in Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran ,
✅ Following our model validation using the existing experimental data, our model suggested critical molecules and candidate medicines involve in surfactant function in rejection of lung transplantation for furtur investigations.
Lung transplantation plays a significant role in improving patients’ quality of life with end-stage lung diseases such as cystic fibrosis, chronic obstructive pulmonary disease, and idiopathic pulmonary fibrosis (1). Lung transplantations have worse outcomes and diagnostic tools for management in comparison to other organ transplants (2). Primary graft dysfunction (PGD) is associated with alveolar damage and is the cause of acute lung injury mortality after lung transplantation (3). Pulmonary surfactant is a lipid and protein complex that makes up the alveolar surface of the lungs. Surfactants have an essential role in lung function, including preventing low-volume alveolar collapse and the formation of pulmonary edema by balancing hydrostatic filtration, maintaining bronchial patency. during normal and forced breathing, and protects the lungs from infection and damage. (4). The deficiency of surfactant function leads to decrease lung compliance, impaired gas exchange, and pulmonary edema. the edematous fluid contains proteins that further reduce the function of the surfactant (5).
Histologic assessment of transbronchial biopsies is complex in lung disease after transplantation. Failure to diagnose the rejection in time may lead to the need for more immunosuppressants (6). Research on the diagnostic and therapeutic applications of microarrays in lung transplantation has been used to define the molecular phenotype of lung rejection and provide clinicians with an accurate biopsy-based system for treating lung transplant recipients. (7). Systems biology science uses various technologies to create a new and promising medical scientific approach, including genomics, proteomics, microbiomics, peptidomics, metabolomics, and antibodyomics. These omics tools generated extensive data of set collections and analyses. The systems biology approach integrates the data obtained by omics tools and diagnostic methods based on clinical phenotyping. These methods promised diagnostic accuracy, a better understanding of pathogenesis, and highly individualized treatments (8).
Since, molecular mechanisms mediating in rejection of lung transplantation related to surfactants are not still comprehensively understood. Therefore, in this in-silico study, we will first analyze the lung transplantation protein-protein interaction network and gene-miRNA-LncRNA regulatory network to identify genes and molecular mechanisms involved in surfactant function in rejection of lung transplantation. Here, we utilize the critical hubs and necessary modules in PPIN and GRN to identify GOs, and functional pathways and repurposing drugs to increase surfactant function for reducing graft rejection in lung disease.
The microarray dataset of lung transplantation was retrieved from the gene expression omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse125478). Using the GSE125478 dataset, Kieran M Halloran et al. investigated the molecular assessment of rejection and damage in lung graft biopsies in lung graft biopsy specimens containing high surfactant and normal surfactant (9). Then, the differentially expressed genes (DEGs) based on log2FC>1.5 or -1.5 fold and P-value<0.01 were separated. This study was confirmed by the ethics committee under the code number IR.SBMU .RETECH.REC.1399.1050. Figure 1 represents the graphical workflow of this study.
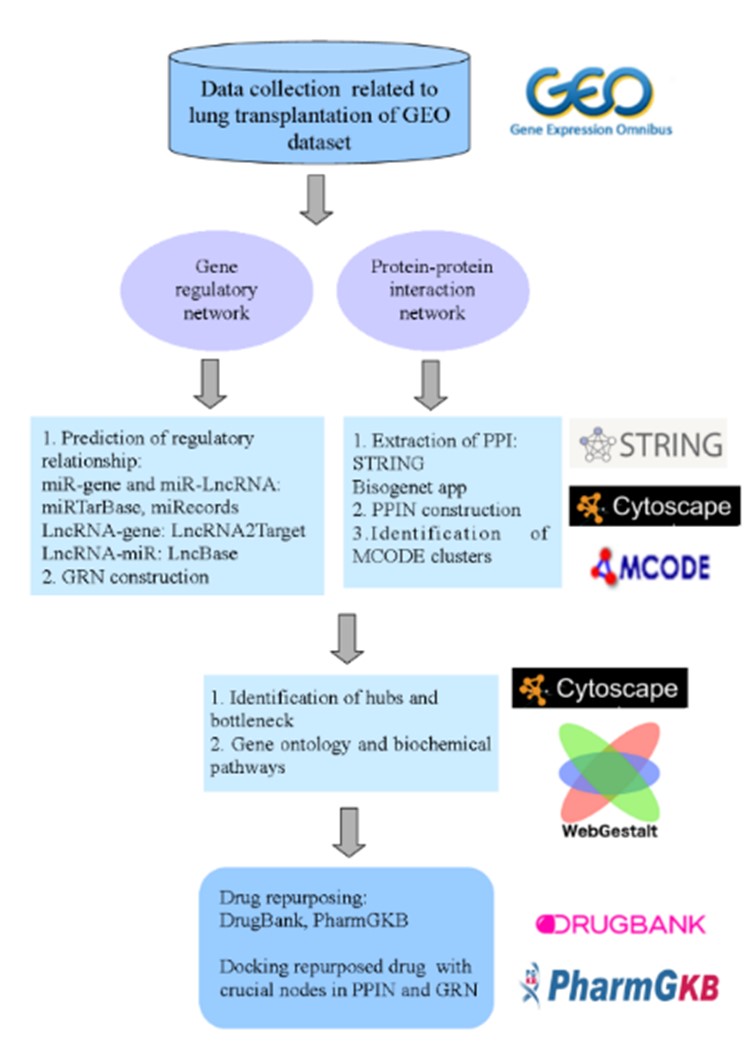
Figure 1. Study workflow
PPIN construction and topological analysis
The DEGs were input into the STRING database (https://string-db.org/) and Bisogenet app (HPRD database) separately to identify the protein-protein interaction network. In the STRING database, protein-protein interactions with a score>0.7 were selected for each gene set. We then merged interactions obtained of STRING and Bisogenet to Cytoscape software (3.5.1) (https://cytoscape.org/). The topological analysis was done by NetworkAnalyzer and identified the top 10% of the nodes having the highest degree and betweenness centrality and closeness centrality. The nodes with the highest degrees are identified as hubs with highly connection proteins, bottlenecks define as nodes with the shortest path. and high closeness centrality nodes are nodes with smaller the total distance of a node to other nodes. Finally, a Venn diagram was used to identify shared nodes between highest degree and betweenness centrality and closeness centrality nodes and further analysis.
Module detection
Molecular Complex Detection (MCODE) was used to examine highly connected network regions (clusters) and identify their seed genes.The interconnected regions with scores>3, and Nodes>10 were selected as significant clusters.
Gene regulatory networks (GRNs)
Identification of microRNAs suppressing DEGs
The experimentally validated miRNA-target genes were collected from miRTarBase (https://bio.tools/mirtarbas) and miRecords (http://c1.accurascience.com/miRecords/).
Identification of LncRNAs suppressing DEGs and microRNAs
LncRNA-DEGs and LncRNA-miRNA interactions were extracted from LncRNA2Target v2.0 (http://123.59.132.21/lncrna2target) and LncBase v2 (www.microrna.gr/LncBase) databases.
microRNAs suppressing LncRNAs
MiR-LncRNA interactions was extracted from miRTarBase and miRecords databases.
GRN construction and topological analysis
The (miR-gene, miR -lncRNA, lncRNA- gene, and lncRNA-miR) regulations were integrated to construct the lncRNA-miRNA-gene regulatory network in Cytoscape software for each up-regulated and down-regulated gene set, separately. We then merged four types of regulations and finally, topological analysis lncRNA-gene-miRNA network done by Network Analyzer app for up-regulated and down-regulated networks, separately.
Gene ontology and functional pathways analysis
The gene ontology (GO) and pathways analysis were performed on critical nodes of the PPI network and clusters using WebGestalt (http://www.webgestalt.org/). Biological processes, molecular functions, cellular components, and functional pathways with p-value<0.05 were determined statistically significant.
Identification of crucial genes and drug repurposing
The gene of hub/bottlenecks of PPI/GRN and MCODE clusters were selected as crucial target genes for drug repurposing. We retrieved the drug-gene interactions among two drug databases, including PharmGKB (https://www.pharmgkb.org/), and Drug Bank database v5.1.6 (https://www.drugbank.ca/). The construction and visualization interaction network between selected genes and their related drugs was performed using Cytoscape software.
Molecular docking
Molecular docking was performed to evaluate hydrogen and non-hydrogen bonds, the potential energy of interaction, and the binding model of repurposed drugs with the binding site of predicted critical genes. The structure of critical genes obtained from the PDB database and the docking studies were performed using Autodock vina v. The most favorable conformations with the lowest free energies (∆G) were selected as final dock conformations.
Gene expression change in lung biopsy of lung transplant patient with low and high surfactant. We extracted the 612 DEGs (175 up-regulated and 437 down-regulated DEGs) based on p-value<0.01 and log2FC <-1.5 or >1.5.
PPIN construction and topological analysis
The PPIN were retrieved among the DEGs by STRING and HPRD (Bisogenet app) databases. We then merged them and visualized PPIN for DEGs using the Cytoscape software. This PPIN includes 632 nodes and 1727 edges. The topological analysis identified nodes with the highest degree, betwenness centrality and closeness centrality. The shared nodes (25 nodes) between 10% degree, 10% betweenness centrality, and 10% closeness centrality identified by Venn diagram, available in. These results include 25 nodes that are available in Table 1.
Table 1. The intersection of 10% degree, 10% betweennesscentrality, and 10% closennesscentrality
| Name of proteins | Degree | Betweennesscentrality | Closennesscentrality |
| DNAI2 | 41 | 0.020408 | 0.31415 |
| DNALI1 | 41 | 0.01715 | 0.31629175 |
| CXCL8 | 37 | 0.073541 | 0.31715653 |
| FN1 | 37 | 0.13989 | 0.33918129 |
| DNAH9 | 37 | 0.015801 | 0.30851064 |
| RSPH1 | 36 | 0.034211 | 0.30306989 |
| FOXJ1 | 35 | 0.155654 | 0.35178165 |
| SPAG6 | 34 | 0.049468 | 0.32289492 |
| RSPH4A | 33 | 0.019956 | 0.31737346 |
| CCDC65 | 29 | 0.035401 | 0.29648562 |
| SPEF2 | 29 | 0.039599 | 0.31245791 |
| HYDIN | 29 | 0.023337 | 0.2925599 |
| SCGB1A1 | 28 | 0.078945 | 0.33967789 |
| ICAM1 | 26 | 0.024407 | 0.29219144 |
| DYX1C1 | 25 | 0.016033 | 0.2955414 |
| EFHC1 | 25 | 0.034258 | 0.32110727 |
| SFTPC | 24 | 0.037141 | 0.32110727 |
| KRT5 | 24 | 0.055832 | 0.31672355 |
| SOX2 | 21 | 0.072241 | 0.3286119 |
| NKX2-1 | 21 | 0.026535 | 0.313302 |
| SFTPB | 19 | 0.023505 | 0.30326797 |
| TEKT1 | 19 | 0.019195 | 0.30346632 |
| ENKUR | 18 | 0.015387 | 0.29404309 |
| SPAG17 | 17 | 0.030172 | 0.30287206 |
| SCGB3A2 | 15 | 0.016998 | 0.29935484 |
MCODE detection
The five sub-networks identified based on nodes>10 and score>3 in PPIN. The identified clusters of PPI network and their seed genes are depicted in Figure 2. he HYDIN, PIH1D3, CXCL8, and LMO7 are as seed genes in clusters.
Functional enrichment analysis
The GO and functional pathways analysis were performed on shared genes obtained of intersection 10% degree, 10% betweenness centrality, and 10% closeness centrality of PPIN and the selected PPI clusters :union: using the Webgestalt database.
The gene ontology on shared genes obtained of intersection 10% degree, 10% betweenness centrality, and 10% closeness centrality are represented microtubule bundle formation, cilium organization, and endoderm development as biological processes involved by shared genes. In molecular function terms, dynein light chain binding and motor activity are as significant terms and cellular components shown the most proteins presented motile cilium, ciliary part, cytoplasmic region, and microtubule-associated complex. The reactome pathway analysis shown the most proteins are involved in Interleukin-4 and Interleukin-13 signaling; Defective CSF2RB causes pulmonary surfactant metabolism dysfunction 5 (SMDP5), defective CSF2RA causes pulmonary surfactant metabolism dysfunction 4 (SMDP4), and diseases associated with surfactant metabolism (based on p-value<0.05).
Microtubule-based process, cell projection assembly, and axoneme assembly were identified as significant terms of the :union: of the selected proteins. Most protein clusters are involved in motor activity and dynein binding as a significant molecular function. Most proteins presented cilium, cytoplasmic region, ciliary part, axoneme, dynein complex, and microtubule-associated complex). The pathway analysis of the :union: of the selected proteins showed that most proteins were involved in surfactant metabolism, immune system, integrin cell-surface interactions, and G alpha signaling.
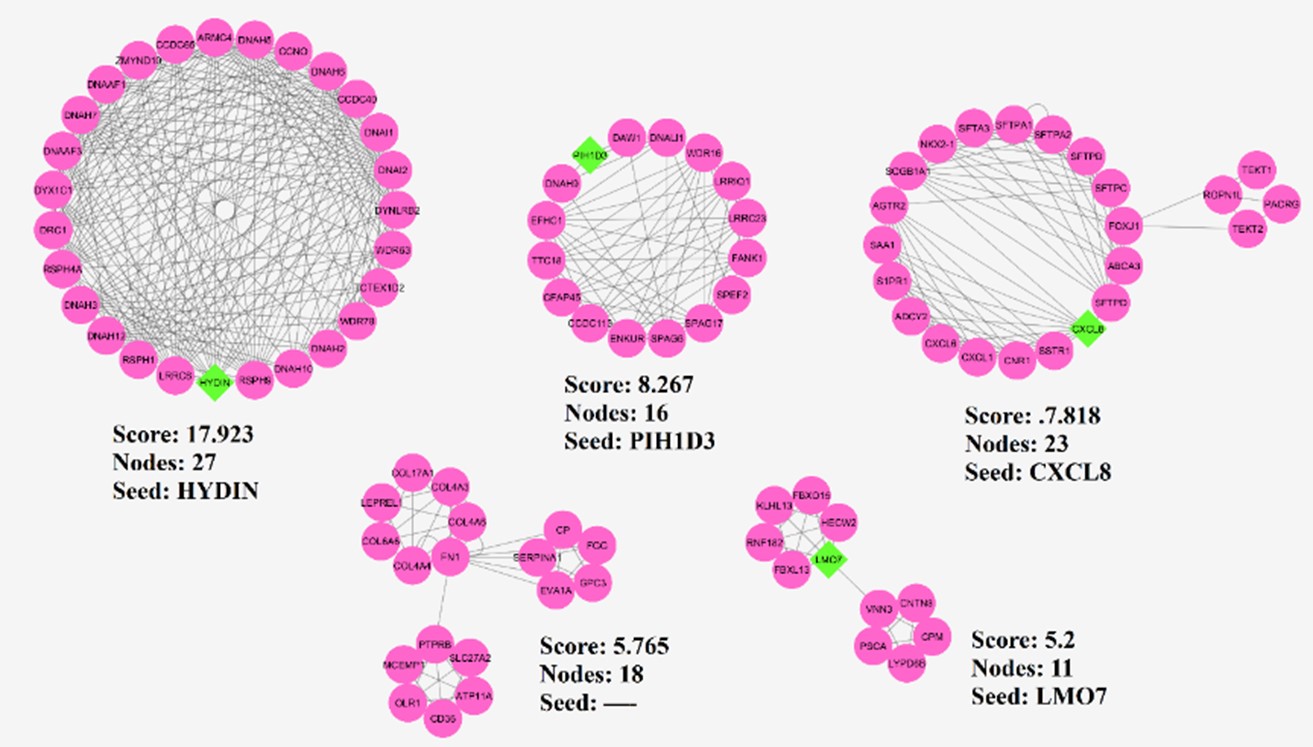
Figure 2. The selected sub-networks of the PPIN of genes studies dataset. Seed genes are shown in green diamonds
Functional enrichment analysis
The GO and functional pathways analysis were performed on shared genes obtained of intersection 10% degree, 10% betweenness centrality, and 10% closeness centrality of PPIN and the selected PPI clusters :union: using the Webgestalt database.
The gene ontology on shared genes obtained of intersection 10% degree, 10% betweenness centrality, and 10% closeness centrality are represented microtubule bundle formation, cilium organization, and endoderm development as biological processes involved by shared genes. In molecular function terms, dynein light chain binding and motor activity are as significant terms and cellular components shown the most proteins presented motile cilium, ciliary part, cytoplasmic region, and microtubule-associated complex. The reactome pathway analysis shown the most proteins are involved in Interleukin-4 and Interleukin-13 signaling; Defective CSF2RB causes pulmonary surfactant metabolism dysfunction 5 (SMDP5), defective CSF2RA causes pulmonary surfactant metabolism dysfunction 4 (SMDP4), and diseases associated with surfactant metabolism (based on p-value<0.05).
Microtubule-based process, cell projection assembly, and axoneme assembly were identified as significant terms of the :union: of the selected proteins. Most protein clusters are involved in motor activity and dynein binding as a significant molecular function. Most proteins presented cilium, cytoplasmic region, ciliary part, axoneme, dynein complex, and microtubule-associated complex). The pathway analysis of the :union: of the selected proteins showed that most proteins were involved in surfactant metabolism, immune system, integrin cell-surface interactions, and G alpha signaling.
GRN construction
The (LncRNA-gene, LncRNA-miRNA, miRNA-LncRNA, and miRNA-gene) regulatory relationships were extracted for each up-regulated and down-regulated gene. These results are present in Table 2. Then, these were constructed the GRN for up-regulated and down-regulated genes separately. The top 5% of nodes with the highest degree, betweenness centrality, and closeness centrality were selected. In up-regulated GRN, SCD is an essential node in the network regulated by GAS5 and MALAT1 LncRNAs. ITGB8, FOXC1, SIX1, FHL2, and TFAP2A are critical nodes in GRN that regulates by MALAT1 and lnrCXCR4 LncRNAs.
Table 2. Summary of four types of regulatory relationships amongLncRNA-gene, LncRNA-miRNA, miRNA-LncRNA, and miRNA-gene interactions.
| Number of pairs | Number of genes | Number of lncRNAs | Number of miRNAs | |
| Up-regulated | ||||
| miR-gene | 2562 | 118 | - | 1075 |
| miR-lncRNA | 915 | - | 83 | 629 |
| lncRNA-miR | 48784 | - | 6030 | 588 |
| lncRNA-gene | 465 | 187 | 45 | - |
| Down-regulated | ||||
| miR-gene | 3736 | 220 | - | 1251 |
| miR-lncRNA | 905 | - | 82 | 622 |
| lncRNA-miR | 76254 | - | 6090 | 624 |
| lncRNA-gene | 1160 | 486 | 52 | - |
Construction of gene-drug interaction network and drug repurposing
The interaction was searched of the medication for 34 critical genes of PPIN and GRN in drug Bbank and pharm GKB databases. For ICAM1 (Hyaluronic acid), FN1 (Zinc, Zinc acetate, Zinc sulfate, Zinc chloride, and Ocriplasmin), KRT5 (Zinc, and Zinc acetate), FHL2 (Gemcitabine), FOXC1 (Oxaliplatin), and CXCL8 (Bevacizumab, Sunitinib, and Crispld2) repurposed interaction drugs.
Molecular docking
To investigate the interaction between repurposed drugs and critical genes, we used Autodock vina v software. The docking results for CXCL8-Sunitinib, FHL2-Gemcitabine, FOXC1-Oxaliplatin, and ICAM1-Hyaluronic acid are represented in Figure 3.
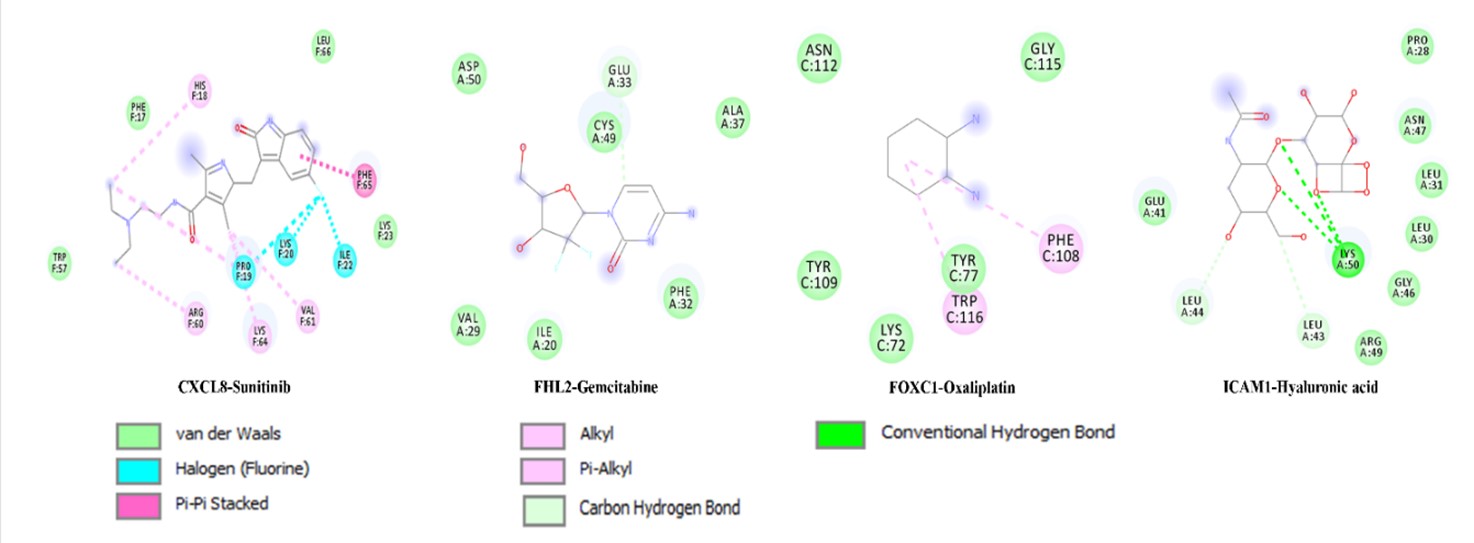
Figure 3. The docking of crucial genes with repurposed new drugs
Discussion
Lung transplantation is therapeutic for patients with end-stage lung disease. The number of lung transplants has increased in worldwide. Despite the post-surgical complications, infections do contribute, and vascular complications are seen transplant rejection (10). Identification of key factor in graft rejection can help to design therapeutic strategies to prevent transplant rejection in lung transplant recipients. Therefore, in this study, we aim to introduce the important factors involved in transplant rejection with bioinformatics analysis.
Our study checked the DEGs related to lung transplantation biopsy in two groups, high surfactant and normal, to investigate critical targets and the molecular mechanisms behind to production and function of surfactants. We constructed protein-protein interaction and lncRNA-miRNA-gene regulatory networks to determine the crucial nodes, gene ontology, and pathways mediating the production and function of surfactants. Finally, hubs and bottlenecks in PPIN and GRN and important seeds were used for repurposing drugs.
This study investigates genes that mediate surfactant production and activity in lung transplant recipients. The most crucial genes included three up-regulated (ICAM1, SCD, and FN1,) and six down-regulated genes (ITGB8, FOXC1, SIX1, FHL2, KRT5, and TFAP2A). On the other hand, the study of gene regulatory networks represented GAS5, MALAT1, and lnrCXCR4 lncRNAs as critical lncRNA regulated hubs and bottlenecks in GRN. Below, we hypothesized that several important genes identified in silico may introduce surfactant function in lung transplant patients.
ICAM1 (intracellular adhesion molecule-1) is a glycoprotein belong to the immunoglobulin superfamily that up-regulated in patients who received lung transplantation with high surfactants. ICAM1 levels increase in inflammatory diseases such as chronic obstructive pulmonary disease, idiopathic pulmonary fibrosis (IPF), and connective tissue disease-associated interstitial pneumonia (CTD-associated IP) (11). Given that the increase in this gene is accompanied by immune responses that may lead to transplant rejection, it should be noted that even in people with an increased surfactant, sufficient attention is paid to pulmonary diseases.
Fibronectin-1 (FN1) and Stearoyl-CoA-Desaturase (SCD) are other up-regulated gene in received lung transplantation patients with high surfactant compared with the normal group. Stearoyl-CoA-Desaturase (SCD) has a key role in lung-specific phospholipids/ surfactant biosynthesis (12).
ITGB8, FOXC1, TFAP2A. SIX1, and FHL2 genes down-regulated in lung transplant receive patient with high surfactant compared with lung transplant receive patient with normal surfactant. The gene regulatory network represents ITGB8 (Integrin Subunit Beta 8) regulated by MALAT1 lncRNA. ITGB8 gene (Integrin Subunit Beta 8) is of the integrin beta chain family that mediates cell-cell and cell-extracellular matrix interactions. It plays a role in the epithelial proliferation of the human airways. ITGB8 has an essential role in activating TGF-B (13). TGF-B is associated with acute and chronic inflammation disorder and is a risk factor for bronchiolitis obliterans syndrome (BOS) and causes primary graft dysfunction (14).
FOXC1, TFAP2A. SIX1, and FHL2 are critical down-regulated genes in the gene regulatory network regulated by lnrCXCR4 lncRNA. Forkhead Box C1 (FOXC1) is a transcription factor that controls proliferation and cellular migration (15). TFAP2A (transcription factor AP-2) is a transcription factor. The role of these genes in surfactant function and mechanisms of lung transplantation rejection not understand.
SIX1 gene (SIX Homeobox 1) is a transcription factor that binds to DNA and controls the activity of other genes. C.Wilson et al. in 2019 reported that expression of SIX1 control pathogenesis of pulmonary fibrosis (16). FHL2 (Four and A Half LIM Domains 2) is a molecular transmitter that regulate transcription (17). Given that surfactant dysfunctions linked to pulmonary fibrosis. Therefore, we suggest reduce expression of SIX1 and FHL2 can help to surfactant function.
Long noncoding RNAs (lncRNAs) are RNAs with lengths exceeding 200 nucleotides that interact with DNA, RNA, and proteins. These molecules act as regulators of protein-coding gene expression by regulation of transcriptional and posttranscriptional processes (18).
The GAS5, MALAT1, and lnrCXCR4 are critical lncRNAs that regulate SCD, ITGB8, FOXC1, SIX1, FHL2, and TFAP2A genes in the up-regulated and down-regulated gene networks. Growth arrest-specific 5 (GAS5) and metastasis KRT5 (Keratin 5) are another down-regulated genes in received lung transplantation patients with high surfactant compared with the normal group. This gene produced by alveolar stem cells could give to type I and type II alveolar cells and finally migrate to the injured area after infection (19). This aberrant differentiation leads to the formation of MUC5B rich honeycomb cysts and pulmonary fibrosis pathogenesis (20). Associated Lung Adenocarcinoma Transcript 1 (MALAT1) lncRNAs regulate the SCD gene. Therefore, the regulation expression of these lncRNAs can moderate the function of the SCD gene.
Our result in gene ontology of selected proteins clusters :union: and intersection between 10% degree, 10% betweenness centrality, and 10% closeness centrality shows the crucial role of the cytoskeleton in transplantation. For instance, microtubule bundle formation is the top biological process at the intersection of 10% degree, 10% betweenness centrality, and 10% closeness centrality. The internal structure of living cells depends on the organization of the cytoskeleton within the cell boundaries. Microtubules are dynamic tubular polymers of the cytoskeleton that interact with various associated proteins. These include multiple types of molecular motors that can form active interconnections between microtubules to form dynamic networks important for the overall organization of the cytoskeleton (21). Some studies show the role of microtubule in surfactant formation (22). Another part of the cytoskeleton which has an essential role in the pulmonary system is cilia. Cilia are microscopic hair-like structures that line the surface of epithelial cells throughout the respiratory system, from the nasal cavity to the terminal bronchioles (23). Cilia beat in concert to protect the airways and lungs from infection and damage and help remove dirt and microbes. After lung transplantation, ciliary beating pattern (CBP) and ciliary frequency (CBF) may be impaired, affecting overall airway clearance and potentially leading to increased respiratory infections (24).
Our result determined that cilia functions such as cilium organization, cilium assembly, and cilium movement are the top biological process in the intersection between the cluster :union: and 10% degree, betweenness centrality and closeness centrality, which illustrate the critical role of cilia in lung transplantation. In addition to driving ciliary beating, axonal dynein binds to double microtubules within the cilia. The outer dynein arm (ODA) and the inner dynein arm (IDA) are two consecutive series of axonal dyneins attached to doublet microtubules (IDA). This is an essential step for cell migration and extracellular fluid flow (25). Our findings reveal that another BP is associated with axoneme assembly and axonemal dynein complex assembly, which explains the involvement of axonemal in distinct regions of cilia and microtubule creation, both of which are important in lung transplantation.
As well as the shared biological processes, some pathways were identified as shared pathways between :union: clusters and 10% degree, betweenness centrality and closeness centrality in Reactome Database. The pathway terms, including Surfactant metabolism and pulmonary surfactant metabolism dysfunction were revealed to be a common critical pathway. Pulmonary surfactant is a complex mixture of phospholipids, neutral lipids, and particular proteins synthesized by type II epithelial cells (AEC2), retained in intracellular organelles called lamellar bodies, and exocytosed to the lungs. Released into the lumen of the alveolus (26). CSF2RA and CSF2RB are two of the primary genes that cause GM-CSF signaling to be blocked by affecting GM-CSF receptor activity. This causes problems with surfactant synthesis and clearance (27). In our study, defective CSF2RB causes pulmonary surfactant metabolism dysfunction 5 (SMDP5) and defective CSF2RA causes pulmonary surfactant metabolism dysfunction 4 (SMDP4) were two pathways that enriched as two significant pathways. In various studies, mutations in two CSF2RA and CSF2RB genes caused the surfactants problems in different disease which related to pulmonary disease. According to the study by Amital et al, during lung transplantation, surfactant instillation improves oxygenation, prevents PGD (Primary graft dysfunction, reduces intubation duration and accelerates the early post-transplantation recovery (28). Our research also found that patients with change surfactant levels have a higher risk of graft rejection.
The other significant pathway in the intersection between 10% degree, betweenness centrality, and closeness centrality was Interleukin-4 and Interleukin-13 signaling. Although IL-4 is essential for tissue repair and counteracting proinflammatory type 1 cytokines, it also contributes to allergic airway inflammation. IL-4, a type 2 cytokine, is also responsible for regulates various human host responses (29). According to a study by Jain-Vora et al., IL-4 synthesis in transgenic mouse airway conducting cells (CCSP-IL-4) also resulted in increased immune cell infiltration, epithelial cell hyperplasia, mucosal metaplasia, collagen staining, and surfactant protein. levels in lung homogenates (30). Our results show Interleukin-4 and Interleukin-13 signaling has a crucial role in lung transplantation.
The results of repurposing drug shown interaction of CXCL8-Sunitinib, FHL2-Gemcitabine, ICAM1-Hyaluronic acid, and FOXC1-Oxaliplatin. CXCL8 (C-X-C Motif Chemokine Ligand 8 or IL8) is a chemokine that is produced by epithelial and endothelial cells, airway smooth muscle cells, and macrophages (31). CXCL8 is a down-regulated factor in received patient transplants with high surfactants compared with those received with typical surfactants. This factor contributes recruitment and activation of neutrophils. CXCL-8 has a crucial role in the phatogenesis of fibrosis, infection, tumor growth, angiogenesis, and arteriosclerosis (32). Sunitinib is a drug interacting with CXCL-8. This drug is an FDA-approved receptor tyrosine kinase inhibitor used to treat renal cell carcinoma (33). Gemcitabine is another repuposed drug that interacted with FHL2. The function of Gemcitabine isn,t clear on FHL2. Therefore, the investigation on this drug can intrude new therapy in patient received lung transplantation. ICAM1 is another up-regulated factor that interacted with hyaluronic acid. The accumulation of hyaluronan activate innate immune and promote allograft rejection (34). Therefore, using this factor can cause transplant rejection in people receiving the transplant. FOXC1 is another crucial molecule that interact with oxaliplatin. Oxaliplatin is a cancer meditation (35). No studies have shown association between oxaliplatin-FOXC1. The study of oxaliplatin could help to preventation of rejection transplantation in received patients.
Conclusion
In this study, the network-based approach (PPIN and GRN) showed the hubs and bottleneck and target genes affecting surfactant production in patients who received a lung transplant. On the other hand, we identified the gene ontology and functional pathways involved in producing surfactants. Microtubule formation, cilium organization, and axoneme assembly were the top critical biological processes involved. Immune response and pathways involved in diseases associated with surfactant metabolism and defective CSF2RB cause pulmonary surfactant metabolism dysfunction Interleukin-4 and Interleukin-13 signaling were pathways active in response to surfactants in patients who received a transplant. The study of a repurposed drug reveals a new drug meditation for the prevention of rejection transplantation. Some of our predictions were verified by other experimental studies, while others still need to be investigated in vitro and in vivo.
Acknowledgements
This study is related to project No.1399/62244 from the Student Research Committee, Shahid Beheshti University of Medical Sciences, Tehran, Iran. We also appreciate the “Student Research Committee” and “Research and Technology Chancellor” in Shahid Beheshti University of Medical Sciences for their financial support of this study.
Funding
This research was funded by Student Research Committee, Shahid Beheshti University of Medical Sciences, Tehran, Iran/ project No.1399/62244.
Authors' contributions
S. A. Mirmotalebisohi, Z. Dehghan, and H. Zali have contributed substantially to the conception or the design of the manuscript, Z. Dehghan, A. Alibakhshi, and F. Yarian to the acquisition, analysis, and interpretation of the data. All authors have participated in drafting the manuscript, and author A revised it critically. All authors read and approved the final version of the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
Received: 2023/08/25 | Accepted: 2023/11/11 | Published: 2024/01/29
| Rights and permissions | |
 |
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License. |